import numpy as np
import matplotlib.pyplot as plt
from epymorph.kit import *
from epymorph.adrio import acs5
from epymorph.adrio import us_tiger
from epymorph.forecasting.pipeline import (
PipelineConfig,
ParticleFilterSimulator,ForecastSimulator,
Observations,
ModelLink,
UnknownParam
)
from epymorph.forecasting.likelihood import Poisson,NegativeBinomial
from epymorph.forecasting.dynamic_params import BrownianMotion,OrnsteinUhlenbeck
from epymorph.forecasting.dynamic_params import GaussianPrior,ExponentialTransform
from epymorph.time import EveryNDays
from epymorph.initializer import RandomLocationsAndRandomSeedForecasting Hospital Admissions Using Particle Filtering
Epymorph’s Bayesian filtering functionality is currently in beta and subject to change.
Scenario
We can use epymorph’s built in particle filter to fit a data set and then run an ensemble forecast from the last data point.
Exercise 1
First, we will develop a multi-node simulation that does not include the effects of movement.
states = ["AZ", "CO", "NM", "NV"]
cutoff_week = 11
full_time_frame = TimeFrame.of("2022-09-15", 7 * 26 + 1)
pf_time_frame = TimeFrame.of("2022-09-15", 7 * cutoff_week)
rume = SingleStrataRUME.build(
ipm=ipm.SIRH(),
mm=mm.No(),
scope=StateScope.in_states(states, year=2015),
init=RandomLocationsAndRandomSeed(num_locations=len(states), seed_max=10_000),
time_frame=pf_time_frame,
params={
"beta": ExponentialTransform("log_beta"),
"gamma": 0.2,
"xi": 1 / 365,
"hospitalization_prob": 200 / 100_000,
"hospitalization_duration": 5.0,
"population": acs5.Population(),
"centroid": us_tiger.InternalPoint(),
},
)
print(f"Last day of inferred data: {pf_time_frame.end_date}")from epymorph.adrio.cdc import InfluenzaStateHospitalizationDaily
adrio = InfluenzaStateHospitalizationDaily(column="admissions")
observations = Observations(
source=adrio,
model_link=ModelLink(
quantity=rume.ipm.select.events("I->H"),
time=rume.time_frame.select.all().group(EveryNDays(1)).agg(),
geo=rume.scope.select.all(),
),
likelihood=NegativeBinomial(r=10),
)The process we choose here is a Black-Karasinski process, a stochastic process whose logarithm is an Ornstein-Uhlenbeck process. The dynamics are controlled by three parameters, the mean, the damping, and the standard deviation. The mean controls the mean of the logarithm of the stationary distribution, the damping controls the rate of mean reversion, and the standard deviation controls the standard deviation of the logarithm of the stationary distribution. We chose the parameters based on experimentation but for many cases parameter estimation algorithms to determine these values may be appropriate.
num_realizations = 500
unknown_params = {
"log_beta": UnknownParam(
prior=GaussianPrior(
mean=np.log(0.25),
standard_deviation=0.2,
),
dynamics=OrnsteinUhlenbeck(
damping=1 / 35,
mean=np.log(0.25),
standard_deviation=0.2,
),
)
}
particle_filter_simulator = ParticleFilterSimulator(
config=PipelineConfig.from_rume(
rume, num_realizations, unknown_params=unknown_params
),
observations=observations,
)rng = np.random.default_rng(0)
particle_filter_output = particle_filter_simulator.run(rng=rng)extend_duration = 28 # Days
forecast_simulator = ForecastSimulator(
PipelineConfig.from_output(particle_filter_output, extend_duration)
)forecast_output = forecast_simulator.run(rng=rng)from math import ceil
n = len(rume.scope.labels)
cols = 2
rows = ceil(n / cols)
fig, axes = plt.subplots(rows, cols, figsize=(6 * cols, 4 * rows), sharex=True)
axes = axes.flatten()
data_date_range = np.arange(0, rume.time_frame.days, 1)
sim_date_range = np.arange(
rume.time_frame.days, rume.time_frame.days + extend_duration, 1
)
total_date_range = np.arange(0, rume.time_frame.days + extend_duration, 1)
tf = TimeFrame.of(rume.time_frame.start_date, 7 * cutoff_week + extend_duration)
real_data_result = adrio.with_context(scope=rume.scope, time_frame = tf).inspect().result
real_data = real_data_result["value"]
for i in range(n):
ax = axes[i]
ax.set_title(f"State {rume.scope.labels[i]}")
ax.set_xlabel("Days")
lower_pf = np.percentile(
particle_filter_output.posterior_values[:, :, i].T, 2.5, axis=0
).squeeze()
upper_pf = np.percentile(
particle_filter_output.posterior_values[:, :, i].T, 97.5, axis=0
).squeeze()
mean_pf = np.median(
particle_filter_output.posterior_values[:, :, i].T, axis=0
).squeeze()
lower_forecast = np.percentile(
forecast_output.events[:, :, i, 1], 2.5, axis=0
).squeeze()
upper_forecast = np.percentile(
forecast_output.events[:, :, i, 1], 97.5, axis=0
).squeeze()
mean_forecast = np.median(forecast_output.events[:, :, i, 1], axis=0).squeeze()
ax.fill_between(
data_date_range, lower_pf, upper_pf, alpha=0.3, label="95% CI of PF"
)
ax.plot(data_date_range, mean_pf, color="black", label="median of PF")
ax.fill_between(
sim_date_range,
lower_forecast,
upper_forecast,
alpha=0.3,
label="95% CI of forecast",
)
ax.plot(sim_date_range, mean_forecast, color="purple", label="median of forecast")
ax.scatter(
total_date_range, real_data[:, i], color="blue", marker="x", label="real data"
)
ax.legend()
for j in range(i + 1, rows * cols):
fig.delaxes(axes[j])
plt.tight_layout()
plt.show()
fig.tight_layout()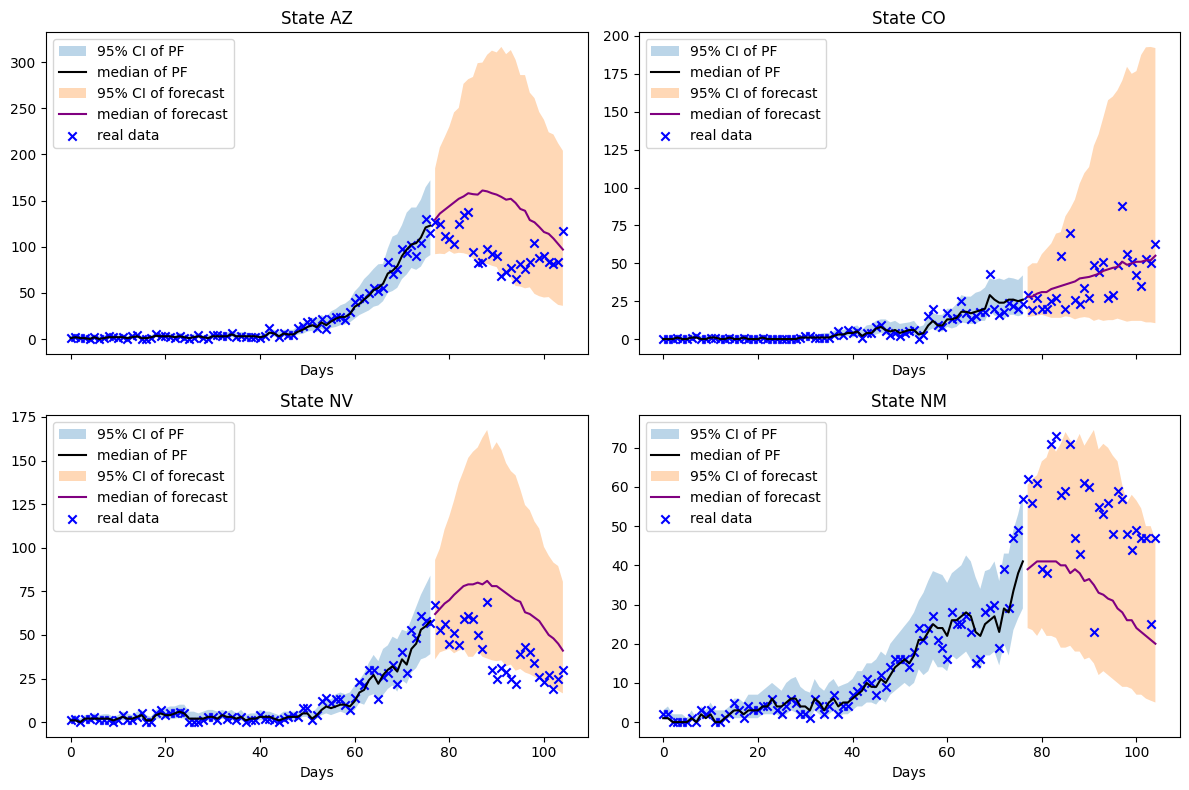
We can layer the particle filter and forecast to incorporate new data points. Let’s examine how the particle filter performs when we incorporate the next week’s data and forecast another four weeks ahead.
extend_pf = 7 # Extend the filter by 7 days
particle_filter_simulator = ParticleFilterSimulator(
config=PipelineConfig.from_output(particle_filter_output, extend_pf),
observations=observations,
)particle_filter_output_update = particle_filter_simulator.run(rng=rng)forecast_simulator = ForecastSimulator(
PipelineConfig.from_output(particle_filter_output_update, extend_duration)
)forecast_output_update = forecast_simulator.run(rng=rng)combined_particle_filter_events = np.concatenate(
(
particle_filter_output.posterior_values,
particle_filter_output_update.posterior_values,
),
axis=0,
)
n = len(rume.scope.labels)
cols = 2
rows = ceil(n / cols)
fig, axes = plt.subplots(rows, cols, figsize=(6 * cols, 4 * rows), sharex=True)
axes = axes.flatten()
data_date_range = np.arange(0, rume.time_frame.days + extend_pf, 1)
sim_date_range = np.arange(
rume.time_frame.days + extend_pf,
rume.time_frame.days + extend_pf + extend_duration,
1,
)
total_date_range = np.arange(0, rume.time_frame.days + extend_duration + extend_pf, 1)
tf = TimeFrame.of(
rume.time_frame.start_date, 7 * cutoff_week + extend_pf + extend_duration
)
real_data_result = adrio.with_context(scope=rume.scope, time_frame=tf).inspect().result
real_data = real_data_result["value"]
for i in range(n):
ax = axes[i]
ax.set_title(f"State {rume.scope.labels[i]}")
ax.set_xlabel("Days")
lower_pf = np.percentile(
combined_particle_filter_events[:, :, i].T, 2.5, axis=0
).squeeze()
upper_pf = np.percentile(
combined_particle_filter_events[:, :, i].T, 97.5, axis=0
).squeeze()
median_pf = np.median(combined_particle_filter_events[:, :, i].T, axis=0).squeeze()
lower_forecast = np.percentile(
forecast_output_update.events[:, :, i, 1], 2.5, axis=0
).squeeze()
upper_forecast = np.percentile(
forecast_output_update.events[:, :, i, 1], 97.5, axis=0
).squeeze()
median_forecast = np.median(
forecast_output_update.events[:, :, i, 1], axis=0
).squeeze()
ax.axvspan(
rume.time_frame.days,
rume.time_frame.days + extend_pf,
alpha=0.5,
color="gray",
label="Additional data",
)
ax.fill_between(
data_date_range, lower_pf, upper_pf, alpha=0.3, label="95% CI of PF"
)
ax.plot(data_date_range, median_pf, color="black", label="median of PF")
ax.fill_between(
sim_date_range,
lower_forecast,
upper_forecast,
alpha=0.3,
label="95% CI of forecast",
)
ax.plot(sim_date_range, median_forecast, color="purple", label="median of forecast")
ax.scatter(
total_date_range, real_data[:, i], color="blue", marker="x", label="real data"
)
ax.legend()
for j in range(i + 1, rows * cols):
fig.delaxes(axes[j])
plt.tight_layout()
plt.show()
fig.tight_layout()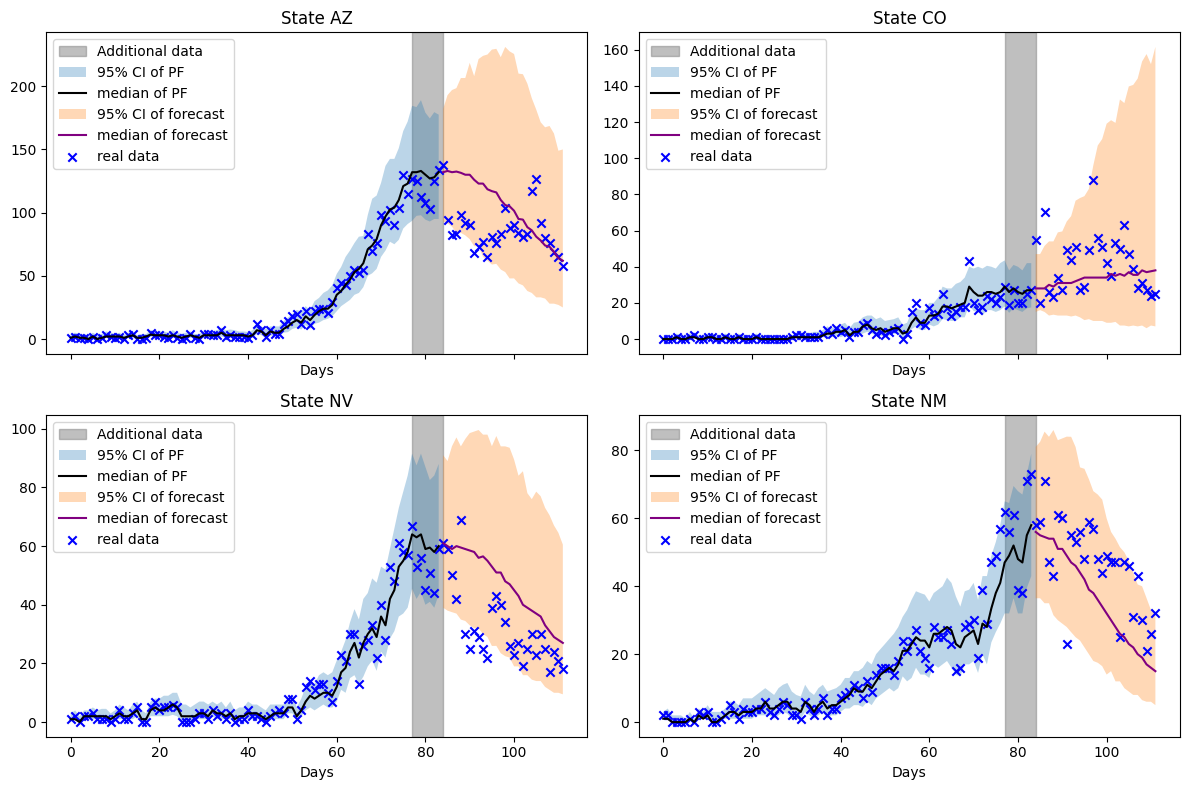
Notes
We see a narrowing of the forecast uncertainty as more data is assimilated by the filter. The downwards trajectory of hospitalizations is captured in 3 of the 4 states. Colorado’s hospitalizations admissions exhibit sub-exponential growth which proves a challenge for this model.