import numpy as np
import matplotlib.pyplot as plt
from epymorph.attribute import NamePattern
from epymorph.kit import *
from epymorph.simulation import Context
from epymorph.adrio import acs5, us_tiger
from epymorph.forecasting.dynamic_params import (
GaussianPrior,
OrnsteinUhlenbeck,
ExponentialTransform,
)
from epymorph.forecasting.pipeline import (
ParticleFilterSimulator,
EnsembleKalmanFilterSimulator,
Observations,
ModelLink,
PipelineConfig,
UnknownParam,
)
from epymorph.forecasting.likelihood import Gaussian
from epymorph.initializer import RandomLocationsAndRandomSeedComparing the Ensemble Kalman Filter and the Particle Filter
Epymorph’s Bayesian filtering functionality is currently in beta and subject to change.
The epymorph package implements multiple techniques for state and parameter estimation. This notebook compares posterior estimates from the ensemble Kalman filter(EnKF) and particle filter(PF), two statistical filtering algorithms. The EnKF is an approximation to the Kalman filter, an algorithm for Bayesian filtering of linear and Gaussian state space models. Empirically, the EnKF performs well for weakly non-linear models by propagating a number of ensemble members through the Kalman update equations. As the EnKF requires a Gaussian likelihood we have also imposed such a likelihood on the PF for comparison.
Exercise 1
We will first generate synthetic data from a multi-node simulation which includes the effects of movement.
scope = CountyScope.in_states(["AZ"], year=2015)
sim_movement_model = mm.Centroids()
sim_ipm = ipm.SIRH()
sim_time_frame = TimeFrame.of("2015-01-01", 26 * 7)
my_rng = np.random.default_rng(0)We now generate the \(\beta_t\) for each node in the simulation. In this example we are generating data for each Arizona county, so we have a \(\beta_t\) for each.
The process we choose here is a Black-Karasinski (BK) process, a stochastic process whose logarithm is an Ornstein-Uhlenbeck process. The dynamics are controlled by three parameters, the mean, the damping, and the standard deviation. The mean controls the mean of the logarithm of the stationary distribution, the damping controls the rate of mean reversion, and the standard deviation controls the standard deviation of the logarithm of the stationary distribution. The constants \(A\),\(M\), and \(C\) are used to define the explicit discretization of the BK process utilized in the loop.
"""Generate a random time dependent beta"""
log_beta_damping = 1 / 35 * np.ones(scope.nodes)
log_beta_mean = np.log(0.4) * np.ones(scope.nodes)
log_beta_standard_deviation = 0.15 * np.ones(scope.nodes)
initial_log_beta = np.log(0.4) * np.ones(scope.nodes)
delta_t = 1.0
A = np.exp(-log_beta_damping * delta_t)
M = log_beta_mean * (np.exp(-log_beta_damping * delta_t) - 1)
C = log_beta_standard_deviation * np.sqrt(1 - np.exp(-2 * log_beta_damping * delta_t))
log_beta = np.zeros(
(
scope.nodes,
sim_time_frame.duration_days,
)
)
log_beta[:, 0] = initial_log_beta
for day in range(1, sim_time_frame.duration_days):
log_beta[:, day] = (
A * log_beta[:, day - 1] - M + C * my_rng.normal(size=(scope.nodes,))
)
beta = np.exp(log_beta)
plt.title("$\\beta$ over Time by Node")
plt.xlabel("Days")
for node in range(scope.nodes):
plt.plot(beta[node, :], label=f"{scope.labels[node]}")
plt.legend()
plt.show()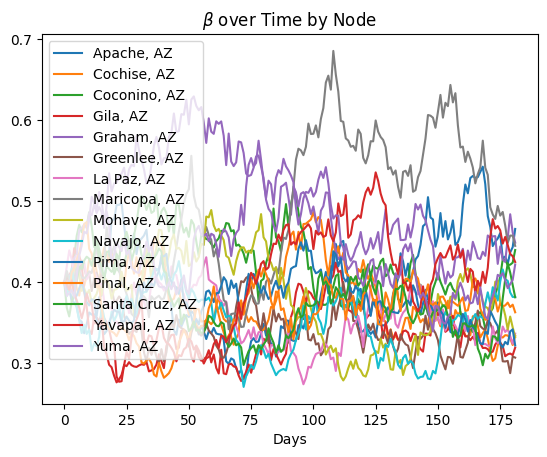
rume = SingleStrataRUME.build(
ipm=sim_ipm,
mm=sim_movement_model,
scope=scope,
init=init.IndexedLocations(selection=np.arange(scope.nodes), seed_size=10_000),
time_frame=sim_time_frame,
params={
"beta": beta.T,
"gamma": 0.1,
"xi": 1 / 90,
"phi": 5,
"hospitalization_prob": 0.05,
"hospitalization_duration": 5,
"centroid": us_tiger.InternalPoint(),
"population": acs5.Population(),
"label": us_tiger.Name(),
},
)sim = BasicSimulator(rume)
with sim_messaging():
out = sim.run(rng_factory=lambda: my_rng)
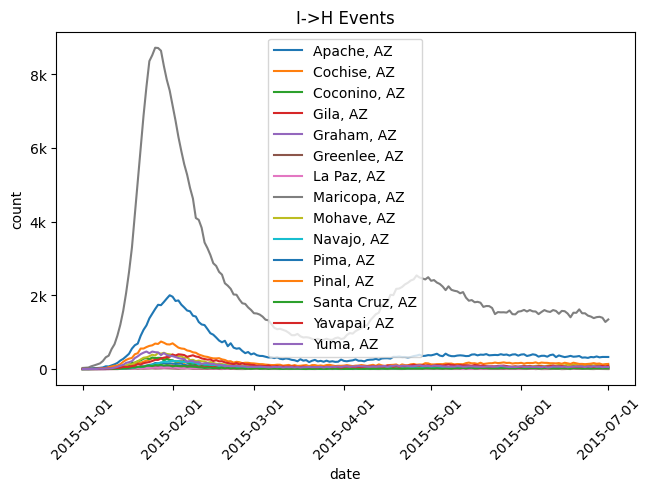
out.plot.line(
geo=rume.scope.select.all(),
time=rume.time_frame.select.all().group("day").agg(),
quantity=rume.ipm.select.events("I->H"),
title="I->H Events",
label_format="{n}",
legend="on",
)
from epymorph.util import to_date_value_array
from epymorph.tools.data import munge
cases_df = munge(
out,
quantity=rume.ipm.select.events("I->H"),
time=rume.time_frame.select.all().group("day").agg(),
geo=rume.scope.select.all(),
)
cases_arr = to_date_value_array(cases_df['time'].to_numpy(),cases_df['I → H'].to_numpy()).reshape(-1,rume.scope.nodes)num_realizations = 100
inf_rume = SingleStrataRUME.build(
ipm=sim_ipm,
mm=sim_movement_model,
scope=scope,
init=RandomLocationsAndRandomSeed(scope.nodes, 50_000),
time_frame=sim_time_frame,
params={
"beta": ExponentialTransform("log_beta"),
"gamma": 0.1,
"xi": 1 / 90,
"phi": 5,
"hospitalization_prob": 0.05,
"hospitalization_duration": 5,
"centroid": us_tiger.InternalPoint(),
"population": acs5.Population(),
"label": us_tiger.Name(),
},
)my_observations = Observations(
source=cases_arr,
model_link=ModelLink(
geo=inf_rume.scope.select.all(),
time=inf_rume.time_frame.select.all().group("day").agg(),
quantity=inf_rume.ipm.select.events("I->H"),
),
likelihood=Gaussian(100.0),
)
my_unknown_params = {
"log_beta": UnknownParam(
prior=GaussianPrior(
mean=log_beta_mean,
standard_deviation=log_beta_standard_deviation,
),
dynamics=OrnsteinUhlenbeck(
damping=log_beta_damping,
mean=log_beta_mean,
standard_deviation=log_beta_standard_deviation,
),
)
}enkf_simulator = EnsembleKalmanFilterSimulator(
config=PipelineConfig.from_rume(
inf_rume, num_realizations, unknown_params=my_unknown_params
),
observations=my_observations,
)
pf_simulator = ParticleFilterSimulator(
config=PipelineConfig.from_rume(
inf_rume, num_realizations, unknown_params=my_unknown_params
),
observations=my_observations,
)enkf_output = enkf_simulator.run(rng=my_rng)
pf_output = pf_simulator.run(rng=my_rng)import math
real_data_result = my_observations.source
real_data = real_data_result["value"]
real_data_dates = real_data_result["date"][:, 0]
t_range = np.arange(inf_rume.time_frame.duration_days)
posterior_obs_enkf = enkf_output.posterior_values
posterior_obs_pf = pf_output.posterior_values
num_nodes = inf_rume.scope.nodes
labels = inf_rume.scope.labels
cols = 3
rows = math.ceil(num_nodes / cols)
fig, axes = plt.subplots(rows, cols, figsize=(cols * 5, rows * 3))
axes = axes.flatten()
plt.suptitle("Data Fit by Node", fontsize=16, y=1.02)
for node in range(num_nodes):
ax = axes[node]
upper_enkf = np.percentile(posterior_obs_enkf[:, :, node].T, 97.5, axis=0)
lower_enkf = np.percentile(posterior_obs_enkf[:, :, node].T, 2.5, axis=0)
data_median_enkf = np.median(posterior_obs_enkf[:, :, node].T, axis=0)
upper_pf = np.percentile(posterior_obs_pf[:, :, node].T, 97.5, axis=0)
lower_pf = np.percentile(posterior_obs_pf[:, :, node].T, 2.5, axis=0)
data_median_pf = np.median(posterior_obs_pf[:, :, node].T, axis=0)
with np.errstate(under="warn"):
rmse_enkf = np.round(
np.sqrt(np.mean((data_median_enkf - real_data[:, node]) ** 2)), 2
)
rmse_pf = np.round(
np.sqrt(np.mean((data_median_pf - real_data[:, node]) ** 2)), 2
)
ax.set_title(
f"{labels[node]} RMSE EnKF: {rmse_enkf}, RMSE PF: {rmse_pf}", fontsize=12
)
ax.set_xlabel("Days")
ax.plot(
t_range,
data_median_enkf,
"--",
color="black",
linewidth=1,
label="median of EnKF",
)
ax.plot(
t_range, data_median_pf, "--", color="orange", linewidth=1, label="median of PF"
)
ax.fill_between(
t_range, lower_enkf, upper_enkf, color="blue", alpha=0.3, label="95% CI EnKF"
)
ax.fill_between(
t_range, lower_pf, upper_pf, color="green", alpha=0.3, label="95% CI PF"
)
ax.scatter(t_range, real_data[:, node], color="red", s=1, marker="x", label="data")
ax.legend(fontsize=8)
for i in range(num_nodes, len(axes)):
axes[i].set_visible(False)
fig.tight_layout(rect=[0, 0, 1, 0.97])
plt.show()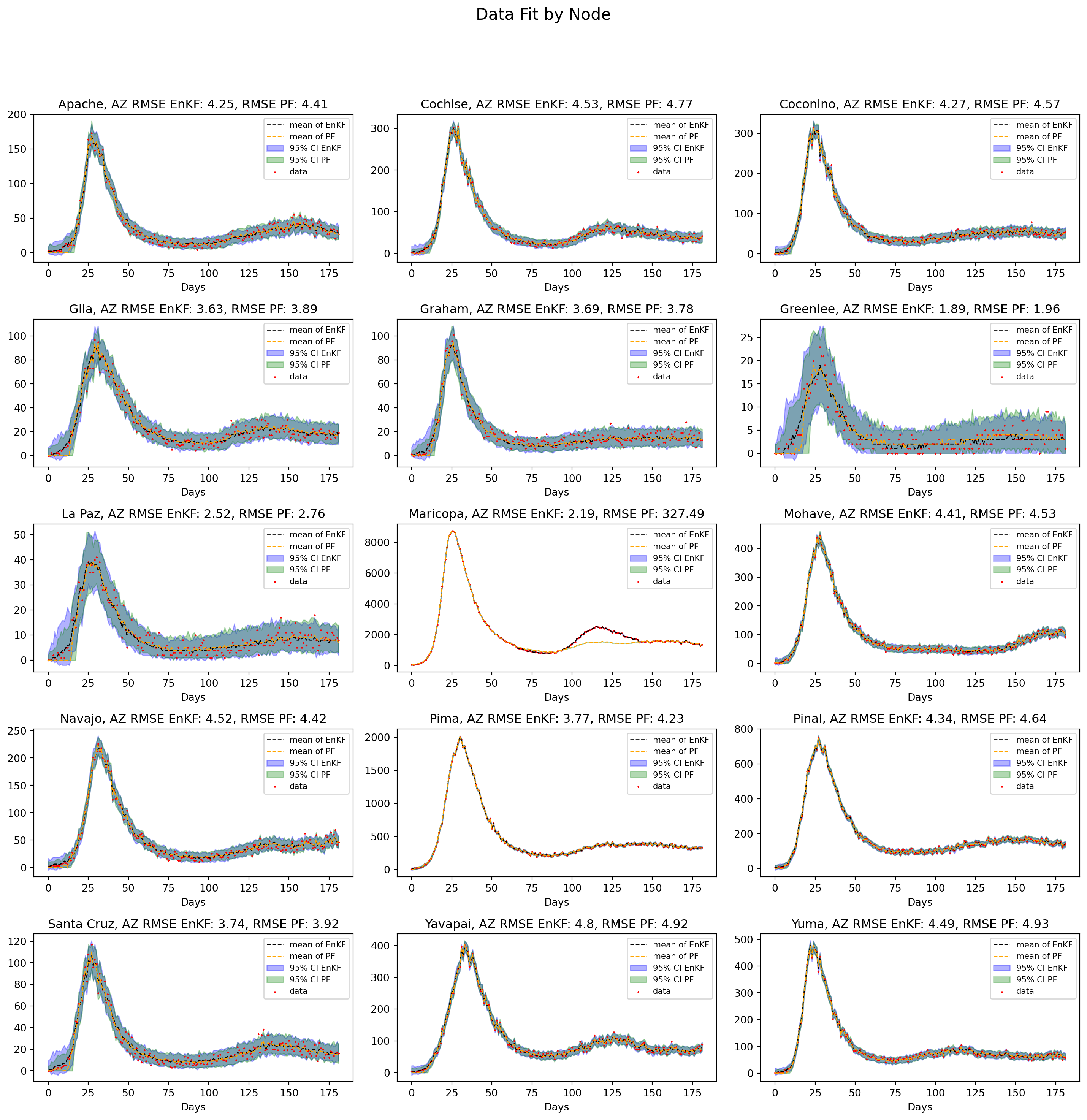
Both the PF and EnKF perform well overall, but the PF has divergence issues for Maricopa county. Due to the finite number of realizations in the PF this divergence can occur for high dimensional systems where PF effectively loses track of the state. The advantage of EnKF in this case is its ability to edit members of the ensemble to match the data, giving it much more flexibility.
num_nodes = inf_rume.scope.nodes
labels = inf_rume.scope.labels
log_beta_enkf = enkf_output.estimated_params[NamePattern.of("log_beta")]
log_beta_pf = pf_output.estimated_params[NamePattern.of("log_beta")]
cols = 3
rows = math.ceil(num_nodes / cols)
fig, axes = plt.subplots(
rows, cols, figsize=(cols * 5, rows * 3), sharex=True, sharey=False
)
axes = axes.flatten()
plt.suptitle(r"$\beta_t$ Fit by Node", fontsize=16, y=1.02)
for node in range(num_nodes):
ax = axes[node]
upper_enkf = np.exp(np.percentile(log_beta_enkf[:, :, node], 97.5, axis=0))
lower_enkf = np.exp(np.percentile(log_beta_enkf[:, :, node], 2.5, axis=0))
beta_median_enkf = np.exp(np.median(log_beta_enkf[:, :, node], axis=0))
upper_pf = np.exp(np.percentile(log_beta_pf[:, :, node], 97.5, axis=0))
lower_pf = np.exp(np.percentile(log_beta_pf[:, :, node], 2.5, axis=0))
beta_median_pf = np.exp(np.median(log_beta_pf[:, :, node], axis=0))
with np.errstate(under="warn"):
rmse_enkf = np.round(
np.sqrt(np.mean((beta_median_enkf - beta[node, :]) ** 2)), 2
)
rmse_pf = np.round(np.sqrt(np.mean((beta_median_pf - beta[node, :]) ** 2)), 2)
ax.set_title(
f"{labels[node]} RMSE EnKF: {rmse_enkf}, RMSE PF: {rmse_pf}", fontsize=12
)
ax.fill_between(
t_range, lower_enkf, upper_enkf, alpha=0.3, color="blue", label="95% CI EnKF"
)
ax.fill_between(
t_range, lower_pf, upper_pf, alpha=0.3, color="green", label="95% CI PF"
)
ax.plot(
t_range,
beta_median_enkf,
"--",
color="black",
linewidth=1,
label="median of EnKF",
)
ax.plot(
t_range, beta_median_pf, "--", color="orange", linewidth=1, label="median of PF"
)
ax.scatter(t_range, beta[node, :], color="red", s=1, marker="x", label="True Value")
ax.legend(fontsize=8)
ax.set_xlabel("Days")
for i in range(num_nodes, len(axes)):
axes[i].set_visible(False)
fig.tight_layout(rect=[0, 0, 1, 0.97])
plt.show()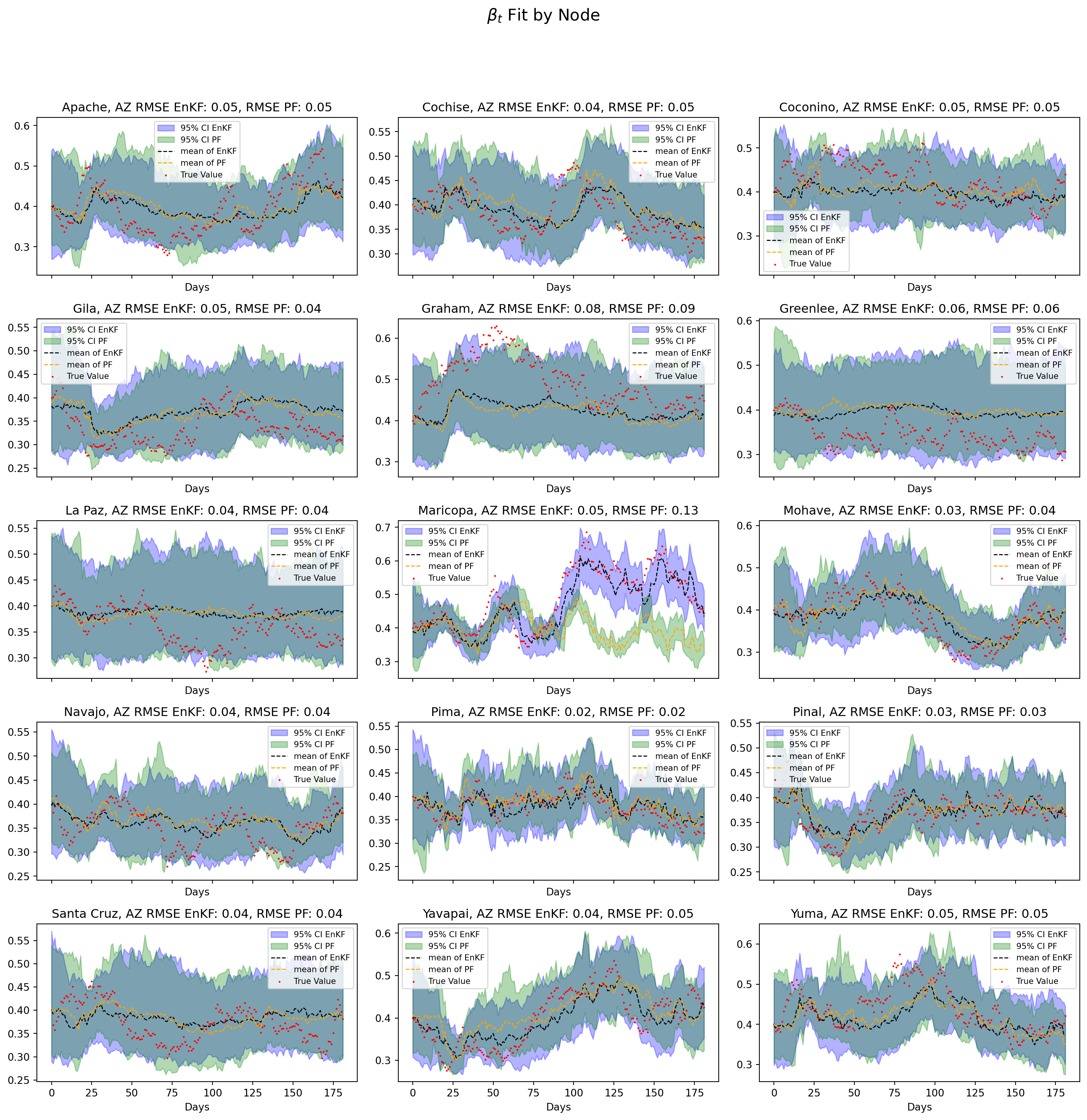
Notes
The divergence of the particle filter for Maricopa county has also manifested in the estimation of \(\beta_t\). The PF fails to capture the upwards trajectory of \(\beta_t\) as opposed to the EnKF which provides a reasonable pathwise approximation.