import numpy as np
import matplotlib.pyplot as plt
from epymorph.kit import *
from epymorph.adrio import acs5
from epymorph.adrio import us_tiger
from epymorph.forecasting.pipeline import (
PipelineConfig,
ParticleFilterSimulator,
Observations,
ModelLink,
UnknownParam,
)
from epymorph.forecasting.likelihood import Poisson
from epymorph.forecasting.dynamic_params import BrownianMotion
from epymorph.forecasting.dynamic_params import GaussianPrior, ExponentialTransform
from epymorph.adrio.cdc import (
InfluenzaStateHospitalization,
InfluenzaStateHospitalizationDaily,
)
from epymorph.time import EveryNDays
from epymorph.initializer import RandomLocationsAndRandomSeedEstimating space- and time-varying transmission rate from data
Epymorph’s Bayesian filtering functionality is currently in beta and subject to change.
Scenario
It is common that we want to account for and quanitfy heterogeneity in transmission rates among localities to better understand the regional picture of pathogen spread. Therefore, we may have a spatial model with multiple nodes, and we want to understand from data how the transmission rate varies among localities. We may also want to quantify this spatial heterogeneity while also accounting for host movement among localities. In epymorph, we have built convenient parameter estimation functionality that allows us to estimate location- and time-varying parameters. In this scenario, we will explore how to set up the model and particle filter simulation for a multi-node system.
Exercise 1
First, we will develop a multi-node simulation that does not include the effects of movement.
states = ["AZ", "CO", "NM", "NV"]
rume = SingleStrataRUME.build(
ipm=ipm.SIRH(),
mm=mm.No(),
scope=StateScope.in_states(states, year=2015),
init=RandomLocationsAndRandomSeed(num_locations=len(states), seed_max=10_000),
time_frame=TimeFrame.of("2022-09-15", 7 * 26 + 1),
params={
"beta": ExponentialTransform("log_beta"),
"gamma": 0.2,
"xi": 1 / 365,
"hospitalization_prob": 200 / 100_000,
"hospitalization_duration": 5.0,
"population": acs5.Population(),
"centroid": us_tiger.InternalPoint(),
},
)observations = Observations(
source=InfluenzaStateHospitalization(),
model_link=ModelLink(
quantity=rume.ipm.select.events("I->H"),
time=rume.time_frame.select.all().group(EveryNDays(7)).agg(),
geo=rume.scope.select.all(),
),
likelihood=Poisson(),
)unknown_params = {
"log_beta": UnknownParam(
prior=GaussianPrior(mean=np.log(0.2), standard_deviation=0.5),
dynamics=BrownianMotion(voliatility=0.1),
)
}num_realizations = 100
particle_filter_simulator = ParticleFilterSimulator(
config=PipelineConfig.from_rume(
rume, num_realizations, unknown_params=unknown_params
),
observations=observations,
)rng = np.random.default_rng(0)
particle_filter_output = particle_filter_simulator.run(rng=rng)from epymorph.attribute import NamePattern
from math import ceil
real_data_result = observations.source.with_context(scope = rume.scope,time_frame = rume.time_frame).inspect().result
real_data = real_data_result["value"]
data_date_range = np.arange(0, rume.time_frame.days, 7)
sim_date_range = np.arange(0, rume.time_frame.days, 1)
n = len(rume.scope.labels)
cols = 2
rows = ceil(n / cols)
fig, axes = plt.subplots(rows, cols, figsize=(6 * cols, 4 * rows))
axes = axes.flatten()
for i in range(n):
ax = axes[i]
ax.set_title(f"State {rume.scope.labels[i]}")
ax.set_xlabel("Days")
lower = np.percentile(
particle_filter_output.posterior_values[:, :, i], 2.5, axis=1
).squeeze()
upper = np.percentile(
particle_filter_output.posterior_values[:, :, i], 97.5, axis=1
).squeeze()
mean = np.median(particle_filter_output.posterior_values[:, :, i], axis=1).squeeze()
ax.fill_between(data_date_range, lower, upper, label="95% CI", alpha=0.3)
ax.plot(data_date_range, mean, color="black", label="median of PF")
ax.scatter(
data_date_range[:-1],
real_data[:, i],
marker="x",
color="red",
label="real data",
)
ax.legend()
for j in range(i + 1, rows * cols):
fig.delaxes(axes[j])
plt.tight_layout()
plt.show()
fig.tight_layout()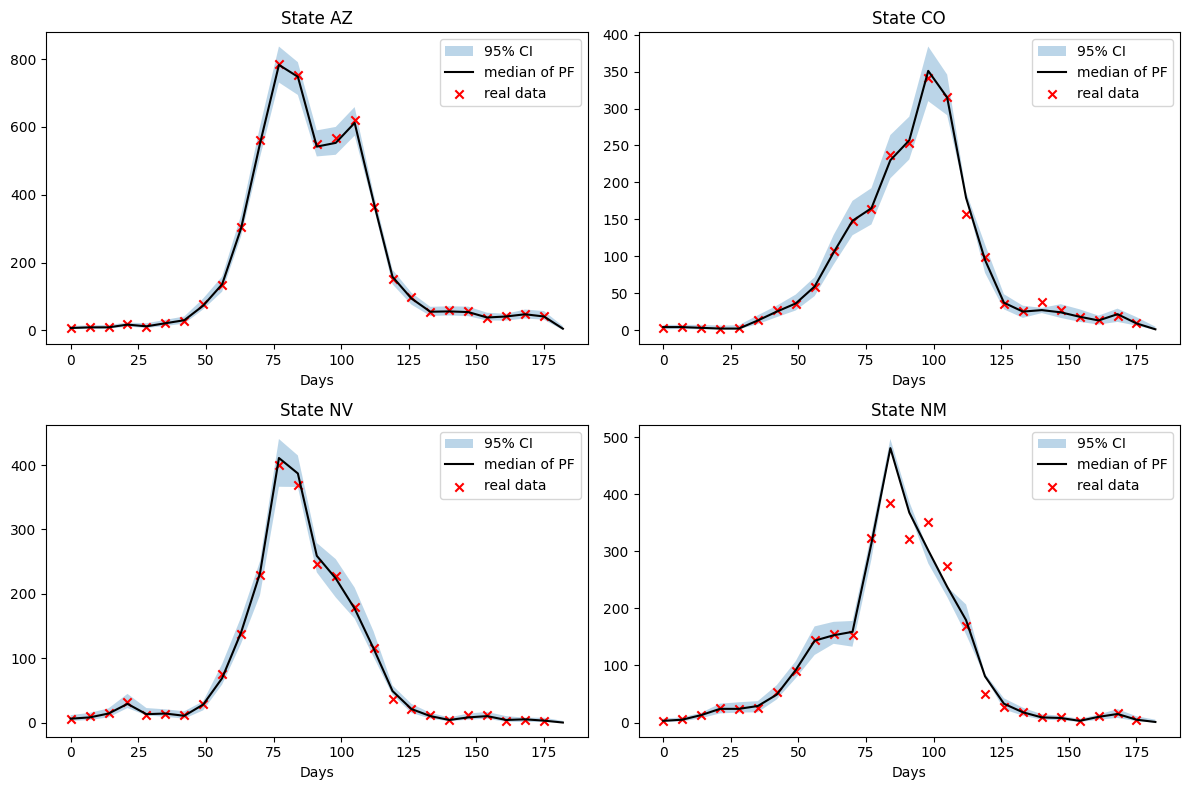
We see a strong fit to the data for three of the states, Nevada has difficulty fitting the model with the data between day 75 and 100. The 95% credible interval describes the interval within which 95% of the particles fall.
cols = 2
rows = ceil(n / cols)
fig, axes = plt.subplots(rows, cols, figsize=(6 * cols, 4 * rows), sharex=True)
axes = axes.flatten()
for i in range(n):
ax = axes[i]
ax.set_title(f"State {rume.scope.labels[i]}")
ax.set_xlabel("Days")
lower = np.percentile(
particle_filter_output.estimated_params[NamePattern.of("log_beta")][:, :, i],
2.5,
axis=0,
).squeeze()
upper = np.percentile(
particle_filter_output.estimated_params[NamePattern.of("log_beta")][:, :, i],
97.5,
axis=0,
).squeeze()
median = np.median(
particle_filter_output.estimated_params[NamePattern.of("log_beta")][:, :, i],
axis=0,
).squeeze()
ax.fill_between(
sim_date_range, np.exp(lower), np.exp(upper), label="95% CI", alpha=0.3
)
ax.plot(sim_date_range, np.exp(median), color="black", label="median of PF")
ax.legend()
for j in range(i + 1, rows * cols):
fig.delaxes(axes[j])
plt.tight_layout()
plt.show()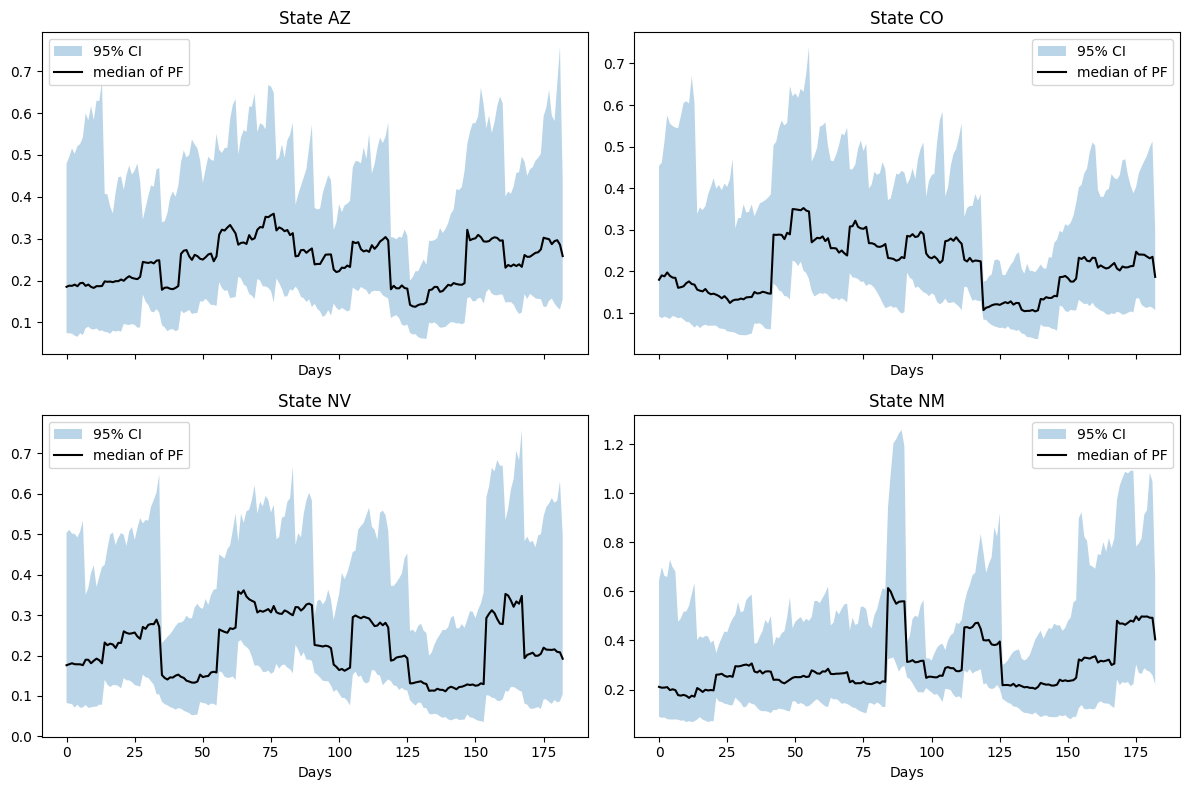
We see a relatively stationary value of \(\beta\) for each location. While seasonal osciallations can be expected in \(\beta\) for certain diseases, agressive and non-smooth oscillations are often indicative of model error.
a = particle_filter_output.compartments.shape[3]
compartment_labels = ["S", "I", "R", "H"]
rows = n
cols = a
fig, axes = plt.subplots(rows, cols, figsize=(6 * cols, 3.5 * rows), sharex=True)
axes = np.atleast_2d(axes)
for i in range(n):
for k in range(a):
ax = axes[i, k]
data = particle_filter_output.compartments[:, :, i, k]
lower = np.percentile(data, 2.5, axis=0).squeeze()
upper = np.percentile(data, 97.5, axis=0).squeeze()
median = np.median(data, axis=0).squeeze()
ax.fill_between(sim_date_range, lower, upper, label="95% CI", alpha=0.3)
ax.plot(sim_date_range, median, color="black", label="median of PF")
if i == 0:
ax.set_title(f"Compartment {compartment_labels[k]}")
if k == 0:
ax.set_ylabel(f"State {states[i]}")
if i == n - 1:
ax.set_xlabel("Days")
ax.legend()
plt.tight_layout()
plt.show()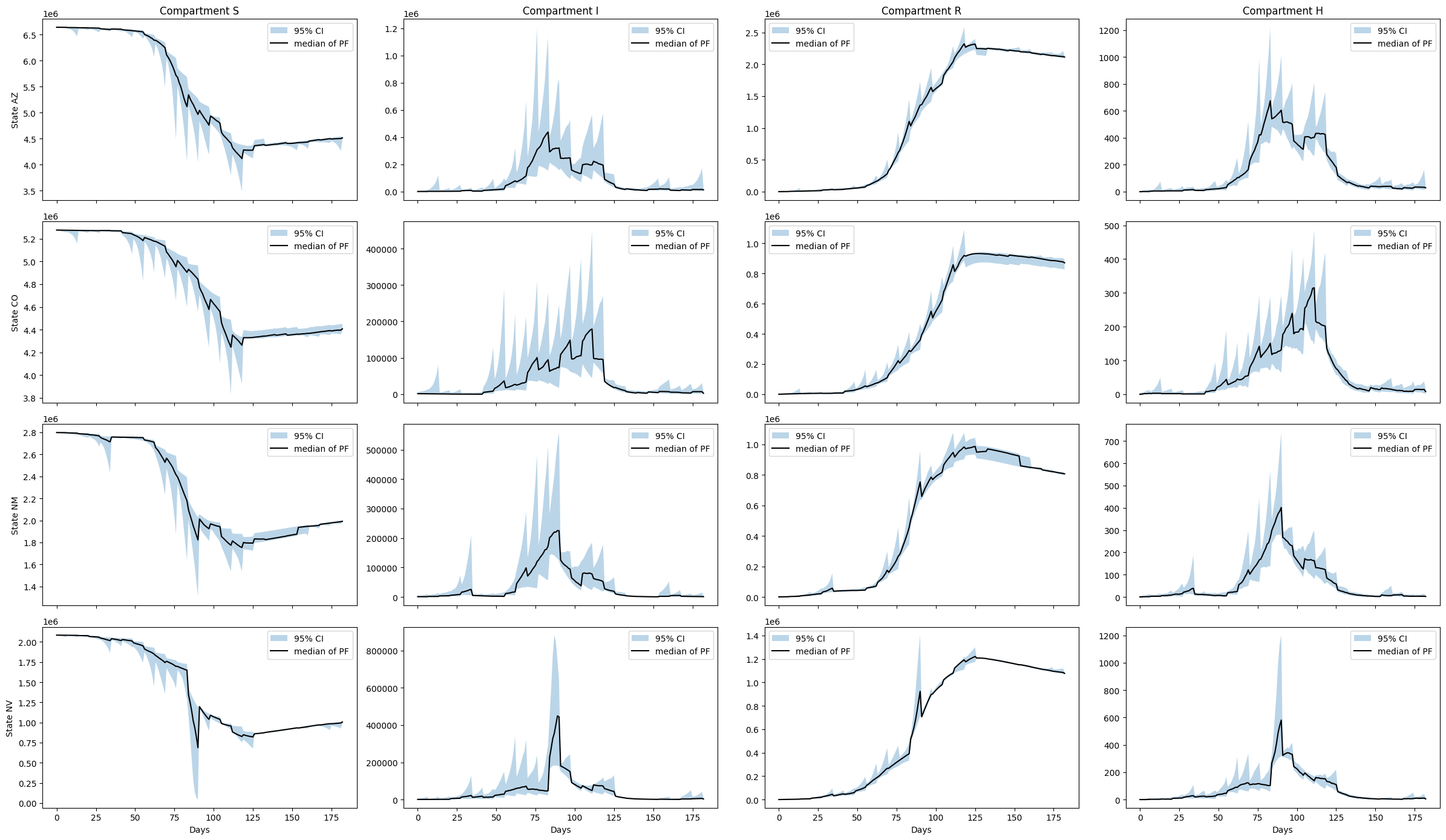
Notes
As in the single node example we observe a strong sawtooth effect due to the sparsity of the data. Posterior distributions adjust sharply each week when new data becomes available. In this example the issue is compounded due to model error. Model error is the discrepancy between the real epidemiological processes that generated the data and the prescribed model. As our SIRH model is a significant approximation of the complex processes underlying influenza spread, the model error is non-trivial.
Exercise 2
The following will fit the same model as above using a daily data source. From August 1st, 2020 to April 30th, 2024 the CDC required hospitals report data on new admissions and total hospitalizations daily. The facility level daily hospitalization data is not available for privacy reasons, however the state aggregations are. We can then fit our model using daily observations of the new hospitalizations and compare the fit to the weekly data.
from epymorph.forecasting.likelihood import NegativeBinomial
adrio_daily = InfluenzaStateHospitalizationDaily(column="admissions")
observations_daily = Observations(
source=adrio_daily,
model_link=ModelLink(
quantity=rume.ipm.select.events("I->H"),
time=rume.time_frame.select.all()
.group(EveryNDays(1))
.agg(), # Changed from 1 day to 7 days
geo=rume.scope.select.all(),
),
likelihood=NegativeBinomial(r=10), # Changed to a negative binomial likelihood
)We now setup a new particle filter simulator with the updated observations and likelihood.
particle_filter_simulator_daily = ParticleFilterSimulator(
config=PipelineConfig.from_rume(
rume, num_realizations, unknown_params=unknown_params
),
observations=observations_daily,
)particle_filter_output_daily = particle_filter_simulator_daily.run(rng=rng)real_data_result = (
observations_daily.source.with_context(scope = rume.scope,time_frame = rume.time_frame).inspect().result
)
real_data = real_data_result["value"]
data_date_range = np.arange(0, rume.time_frame.days, 1)
sim_date_range = np.arange(0, rume.time_frame.days, 1)
n = len(rume.scope.labels)
cols = 2
rows = ceil(n / cols)
fig, axes = plt.subplots(rows, cols, figsize=(6 * cols, 4 * rows), sharex=True)
axes = axes.flatten()
for i in range(n):
ax = axes[i]
ax.set_title(f"State {rume.scope.labels[i]}")
ax.set_xlabel("Days")
lower = np.percentile(
particle_filter_output_daily.posterior_values[:, :, i], 2.5, axis=1
).squeeze()
upper = np.percentile(
particle_filter_output_daily.posterior_values[:, :, i], 97.5, axis=1
).squeeze()
median = np.median(
particle_filter_output_daily.posterior_values[:, :, i], axis=1
).squeeze()
ax.fill_between(data_date_range, lower, upper, alpha=0.3, label="95% CI")
ax.plot(data_date_range, median, color="black", label="Mean of PF")
ax.scatter(
data_date_range, real_data[:, i], marker="x", color="red", label="real data"
)
ax.legend()
for j in range(i + 1, rows * cols):
fig.delaxes(axes[j])
plt.tight_layout()
plt.show()
fig.tight_layout()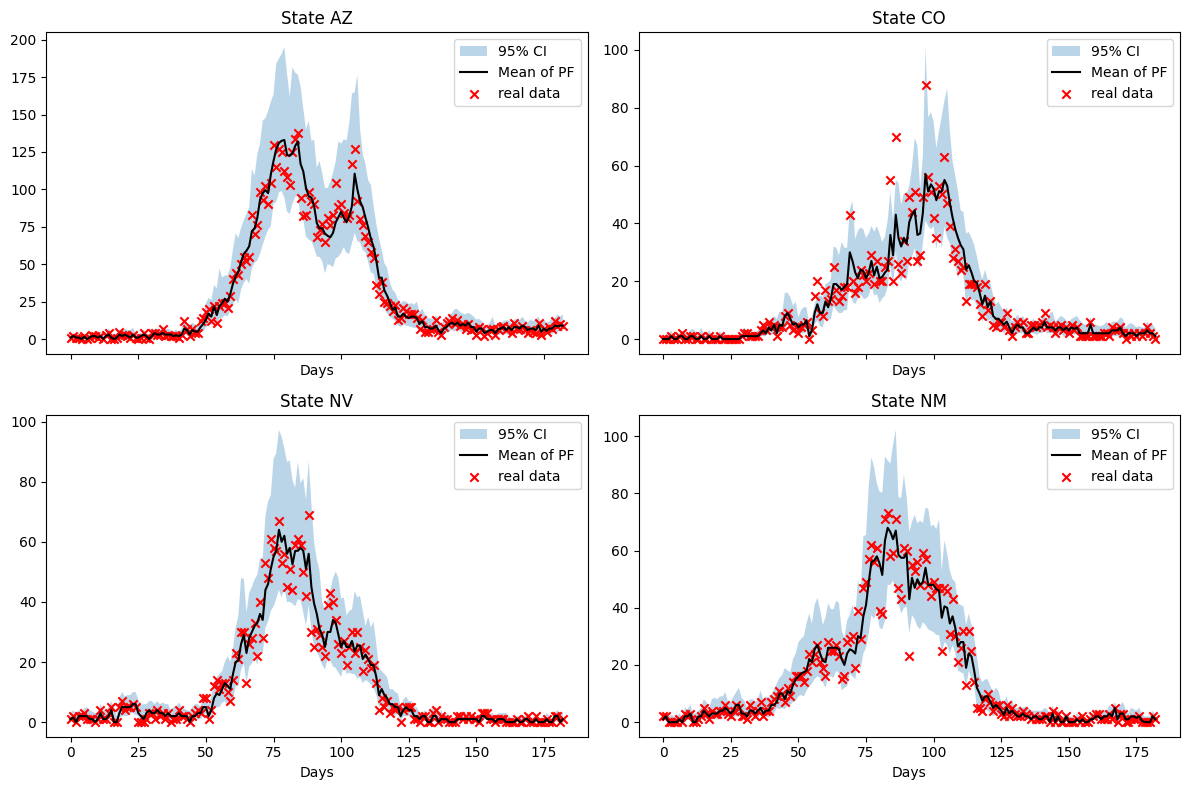
cols = 2
rows = ceil(n / cols)
fig, axes = plt.subplots(rows, cols, figsize=(6 * cols, 4 * rows), sharex=True)
axes = axes.flatten()
for i in range(n):
ax = axes[i]
ax.set_title(f"State {rume.scope.labels[i]}")
ax.set_xlabel("Days")
lower = np.percentile(
particle_filter_output.estimated_params[NamePattern.of("log_beta")][:, :, i],
2.5,
axis=0,
).squeeze()
upper = np.percentile(
particle_filter_output.estimated_params[NamePattern.of("log_beta")][:, :, i],
97.5,
axis=0,
).squeeze()
median = np.median(
particle_filter_output.estimated_params[NamePattern.of("log_beta")][:, :, i],
axis=0,
).squeeze()
lower_daily = np.percentile(
particle_filter_output_daily.estimated_params[NamePattern.of("log_beta")][
:, :, i
],
2.5,
axis=0,
).squeeze()
upper_daily = np.percentile(
particle_filter_output_daily.estimated_params[NamePattern.of("log_beta")][
:, :, i
],
97.5,
axis=0,
).squeeze()
median_daily = np.median(
particle_filter_output_daily.estimated_params[NamePattern.of("log_beta")][
:, :, i
],
axis=0,
).squeeze()
ax.fill_between(
sim_date_range,
np.exp(lower_daily),
np.exp(upper_daily),
alpha=0.3,
label="95% CI for Daily PF",
)
ax.plot(
sim_date_range, np.exp(median_daily), color="black", label="Median of Daily PF"
)
ax.plot(sim_date_range, np.exp(median), label="Median of Weekly PF", color="red")
ax.legend()
for j in range(i + 1, rows * cols):
fig.delaxes(axes[j])
plt.tight_layout()
plt.show()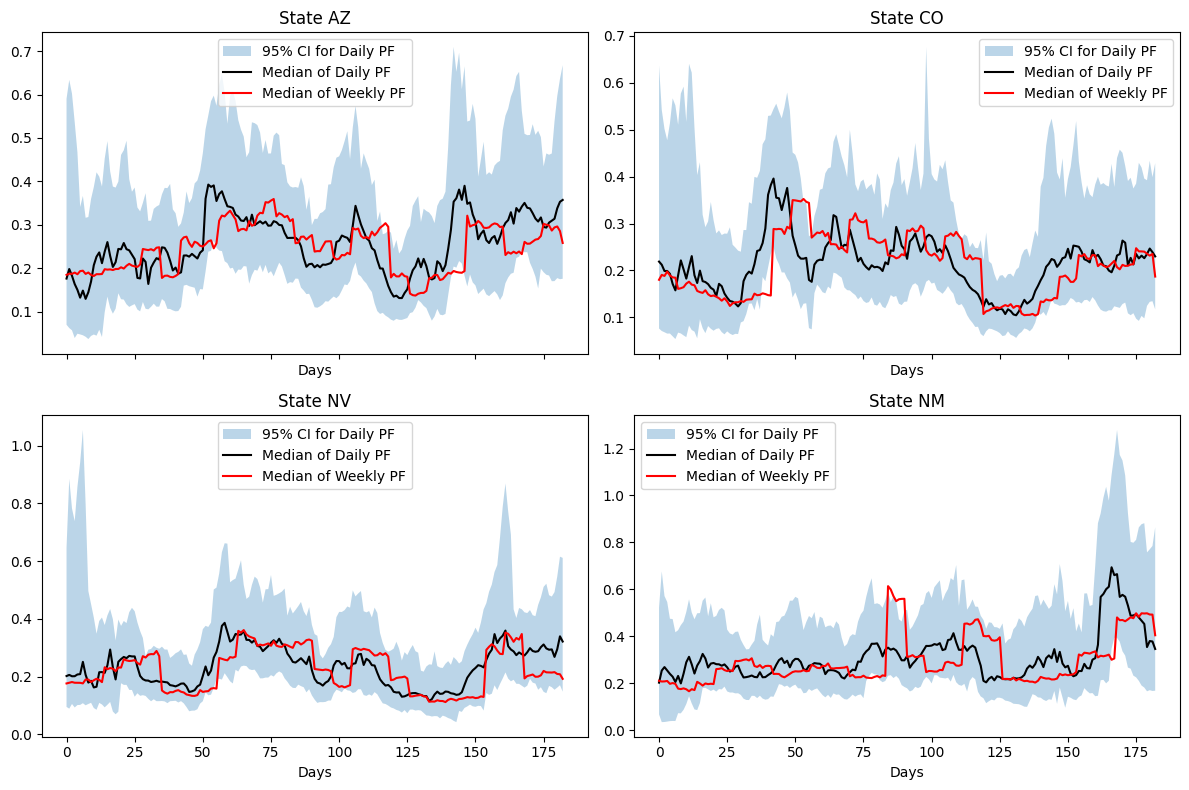
We observe a \(\beta\) which is qualitatively similar to the estimate from the weekly inference. However, the dynamics are more finely resolved and we reduce the stepping effect observed with the weekly estimate.
a = particle_filter_output_daily.compartments.shape[3]
compartment_labels = ["S", "I", "R", "H"]
rows = n
cols = a
fig, axes = plt.subplots(rows, cols, figsize=(6 * cols, 3.5 * rows), sharex=True)
axes = np.atleast_2d(axes)
observations_daily_hosp = InfluenzaStateHospitalizationDaily(column="hospitalizations")
real_data_result_hosp = (
observations_daily_hosp.with_context(scope = rume.scope,time_frame = rume.time_frame).inspect().result
)
real_data_hosp = real_data_result_hosp["value"]
for i in range(n):
for k in range(a):
ax = axes[i, k]
ax.set_xlabel("Days")
data = particle_filter_output_daily.compartments[:, :, i, k]
lower = np.percentile(data, 2.5, axis=0).squeeze()
upper = np.percentile(data, 97.5, axis=0).squeeze()
median = np.median(data, axis=0).squeeze()
ax.fill_between(sim_date_range, lower, upper, alpha=0.3, label="95% CI")
ax.plot(sim_date_range, median, color="black", label="Median of PF")
if k == 3:
ax.scatter(
sim_date_range,
real_data_hosp[:, i],
color="red",
marker="x",
label="Hospitalization data",
)
if i == 0:
ax.set_title(f"Compartment {compartment_labels[k]}")
if k == 0:
ax.set_ylabel(f"State {rume.scope.labels[i]}")
ax.legend()
plt.tight_layout()
plt.show()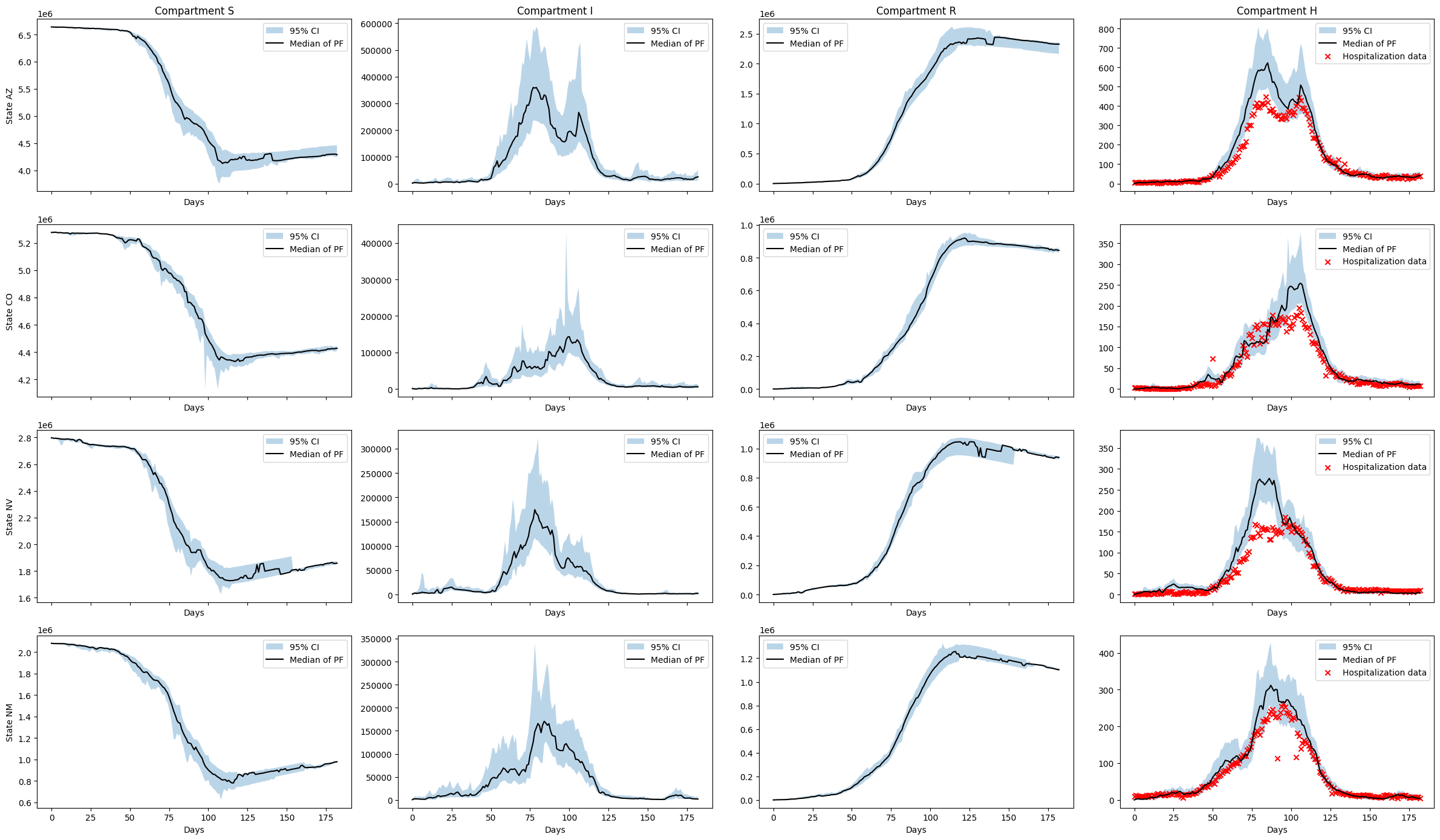
The latent state estimation is vastly improved and the estimated trajectories are reasonable for the \(H\) compartment. The algorithm is more able to account for model error at this daily data resolution.