from epymorph.adrio import acs5, csv
from epymorph.data.ipm.sirh import SIRH
from epymorph.data.mm.no import No
from epymorph.initializer import SingleLocation
import numpy as np
from epymorph.kit import *Estimating time-varying transmission rate from data
Epymorph’s Bayesian filtering functionality is currently in beta and subject to change.
Scenario
A particle filter is an algorithm which can use data to estimate the states and parameters of an epymorph model. The particle filter accomplishes this by combining the data with a mathematical model.
This vignette will demonstrate how to use epymorph’s built-in particle filter to estimate both the time-varying transmission rate (beta) and state variables \(S,I,R,H\) using weekly influenza hospitalization data. In the first exercise, we will define our model setup and generate synthetic data (fake data used for testing) which will be used to validate the particle filter. In the second exercise, we will setup the particle filter and run it on the synthetic data we generated in the first exercise. By validating the particle filter setup on synthetic data first, we can diagnose any issues before moving on to real data. In the third exercise, we will run the particle filter on real influenza hospitalization data. Because we will have already tested the particle filter on synthetic data, this step will be as easy as swapping out the synthetic data for the real data.
Exercise 1
In this first exercise, we will setup our mathematical model and generate synthetic data.
num_weeks = 26
duration = 7 * num_weeks + 1
t = np.arange(0, duration)
true_beta = 0.1 * np.cos(t * 2 * np.pi / (365)) + 0.25import matplotlib.pyplot as plt
plt.plot(t, true_beta)
plt.ylabel("$\\beta$")
plt.xlabel("Days")
plt.title("The transmission rate ($\\beta$) over time")
plt.show()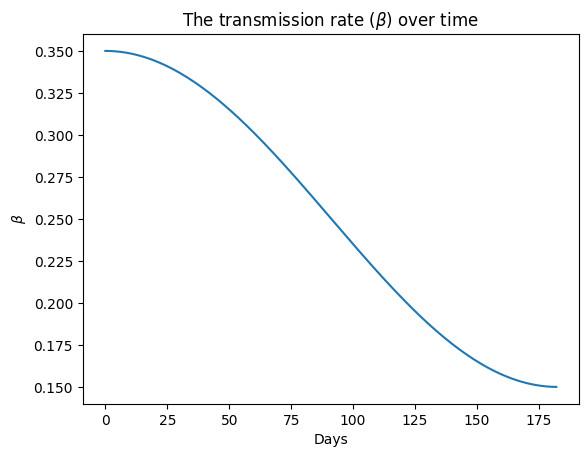
my_ipm = SIRH()
my_ipm.diagram()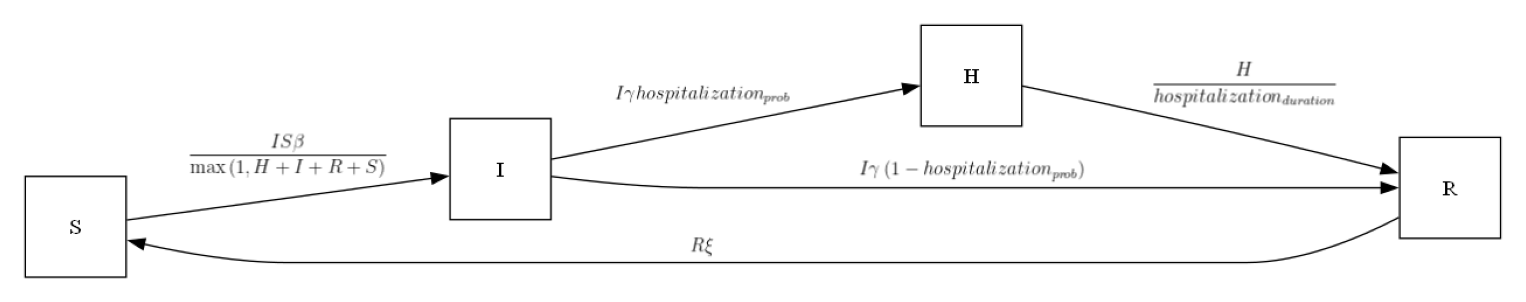
rume = SingleStrataRUME.build(
ipm=SIRH(),
mm=No(),
scope=StateScope.in_states(["AZ"], year=2015),
init=SingleLocation(0,100),
time_frame=TimeFrame.of("2022-09-15", 7 * 26 + 1),
params={
"beta": true_beta,
"gamma": 0.2,
"xi": 1 / 365,
"hospitalization_prob": 200 / 100_000,
"hospitalization_duration": 5.0,
"population": acs5.Population(),
},
)rng = np.random.default_rng(seed=1)
sim = BasicSimulator(rume)
with sim_messaging(live = False):
out = sim.run(rng_factory=(lambda: rng))from epymorph.time import EveryNDays
from epymorph.tools.data import munge
from epymorph.util import to_date_value_array
quantity_selection = rume.ipm.select.events("I->H")
time_selection = rume.time_frame.select.all().group(EveryNDays(7)).agg()
geo_selection = rume.scope.select.all()
cases_df = munge(
out,
quantity=quantity_selection,
time=time_selection,
geo=geo_selection,
)
cases_arr = to_date_value_array(
cases_df["time"].to_numpy(), cases_df["I → H"].to_numpy()
)[...,np.newaxis]out.plot.line(geo = geo_selection,
time = time_selection,
quantity = quantity_selection,
title = 'New Hospitalizations in Arizona',
label_format = "{n}")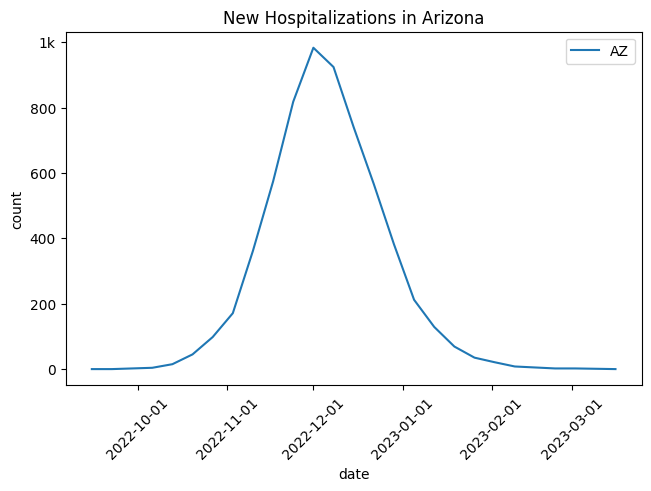
Exercise 2
In this second exercise, we will run the particle filter on the synthetic data we generated and compare the estimate of beta produced by the particle filter to the true values of beta which were used to generate the data.
We now define the rume associated with the particle filter. This represents the model we plan to use for inference. Note that the only difference from the model used for generating the synthetic data is the specification of \(\beta\). We choose to estimate \(\ln \beta\) and perform an exponential transform to ensure \(\beta\) remains positive (a negative \(\beta\) is non-physical).
from epymorph.forecasting.dynamic_params import ExponentialTransform
pf_rume = SingleStrataRUME.build(
ipm=SIRH(),
mm=No(),
scope=StateScope.in_states(["AZ"], year=2015),
init=SingleLocation(0,100),
time_frame=TimeFrame.of("2022-09-15", 7 * 26 + 1),
params={
"beta": ExponentialTransform('log_beta'),
"gamma": 0.2,
"xi": 1 / 365,
"hospitalization_prob": 200 / 100_000,
"hospitalization_duration": 5.0,
"population": acs5.Population(),
},
)from epymorph.forecasting.pipeline import ModelLink
#Make sure the selectors reference the pf_rume not the rume for the synthetic data.
quantity_selection = pf_rume.ipm.select.events("I->H")
time_selection = pf_rume.time_frame.select.all().group(EveryNDays(7)).agg()
geo_selection = pf_rume.scope.select.all()
model_link=ModelLink(
quantity=quantity_selection,
time=time_selection,
geo=geo_selection,
)from epymorph.forecasting.likelihood import Poisson
from epymorph.forecasting.pipeline import Observations
from epymorph.adrio.adrio import Context
observations = Observations(
source=cases_arr,
model_link=model_link,
likelihood=Poisson(),
)from epymorph.forecasting.dynamic_params import GaussianPrior,BrownianMotion
from epymorph.forecasting.pipeline import UnknownParam
my_unknown_params = {
"log_beta": UnknownParam(
prior=GaussianPrior(
mean=np.log(0.4),
standard_deviation=0.2
),
dynamics=BrownianMotion(
voliatility = 0.07
),
)
}from epymorph.forecasting.pipeline import PipelineConfig, ParticleFilterSimulator
num_realizations = 100
particle_filter_simulator = ParticleFilterSimulator(
config=PipelineConfig.from_rume(
pf_rume, num_realizations, unknown_params=my_unknown_params
),
observations=observations,
)particle_filter_output = particle_filter_simulator.run(rng=rng)The particle filter output emcompasses a number of useful metrics and data outputs from the run. Here, we are interested in the estimates of \(\beta,S,I,R,H\) and associated 95% credible interval. We will also plot the fit to the data, this provides a useful sanity check to ensure the algorithm is working appropriately.
from epymorph.simulation import Context
from epymorph.attribute import NamePattern
context = Context.of(scope=pf_rume.scope, time_frame=pf_rume.time_frame)
data_date_range = np.arange(0, pf_rume.time_frame.days, 7)
sim_date_range = np.arange(0, pf_rume.time_frame.days, 1)
fig, axes = plt.subplots(3, 2, figsize=(12, 10), sharex=True)
axes[0, 0].set_title("Fit to the Data")
axes[0, 0].fill_between(
data_date_range,
np.percentile(particle_filter_output.posterior_values, 2.5, axis=1).squeeze(),
np.percentile(particle_filter_output.posterior_values, 97.5, axis=1).squeeze(),
label="95% CI",
alpha=0.3,
)
axes[0, 0].plot(
data_date_range,
np.median(particle_filter_output.posterior_values, axis=1).squeeze(),
color="black",
label="median of PF",
)
axes[0, 0].scatter(
data_date_range, cases_df["I → H"], marker="x", color="red", label="Data points"
)
axes[0, 0].legend()
axes[0, 1].set_title("Fit to the S Compartment")
axes[0, 1].fill_between(
sim_date_range,
np.percentile(
particle_filter_output.compartments[:, :, 0, 0], 2.5, axis=0
).squeeze(),
np.percentile(
particle_filter_output.compartments[:, :, 0, 0], 97.5, axis=0
).squeeze(),
label="95% CI",
alpha=0.3,
)
axes[0, 1].plot(sim_date_range, out.dataframe["S"], color="red", label="Data points")
axes[0, 1].plot(
sim_date_range,
np.median(particle_filter_output.compartments[:, :, 0, 0], axis=0).squeeze(),
color="black",
label="median of PF",
)
axes[0, 1].legend()
axes[1, 0].set_title("Fit to the I Compartment")
axes[1, 0].fill_between(
sim_date_range,
np.percentile(
particle_filter_output.compartments[:, :, 0, 1], 2.5, axis=0
).squeeze(),
np.percentile(
particle_filter_output.compartments[:, :, 0, 1], 97.5, axis=0
).squeeze(),
label="95% CI",
alpha=0.3,
)
axes[1, 0].plot(sim_date_range, out.dataframe["I"], color="red", label="Data points")
axes[1, 0].plot(
sim_date_range,
np.median(particle_filter_output.compartments[:, :, 0, 1], axis=0).squeeze(),
color="black",
label="median of PF",
)
axes[1, 0].legend()
axes[1, 1].set_title("Fit to the R Compartment")
axes[1, 1].fill_between(
sim_date_range,
np.percentile(
particle_filter_output.compartments[:, :, 0, 2], 2.5, axis=0
).squeeze(),
np.percentile(
particle_filter_output.compartments[:, :, 0, 2], 97.5, axis=0
).squeeze(),
label="95% CI",
alpha=0.3,
)
axes[1, 1].plot(sim_date_range, out.dataframe["R"], color="red", label="Data points")
axes[1, 1].plot(
sim_date_range,
np.median(particle_filter_output.compartments[:, :, 0, 2], axis=0).squeeze(),
color="black",
label="median of PF",
)
axes[1, 1].legend()
axes[2, 0].set_title("Fit to the H Compartment")
axes[2, 0].set_xlabel("Days")
axes[2, 0].fill_between(
sim_date_range,
np.percentile(
particle_filter_output.compartments[:, :, 0, 3], 2.5, axis=0
).squeeze(),
np.percentile(
particle_filter_output.compartments[:, :, 0, 3], 97.5, axis=0
).squeeze(),
label="95% CI",
alpha=0.3,
)
axes[2, 0].plot(sim_date_range, out.dataframe["H"], color="red", label="Data points")
axes[2, 0].plot(
sim_date_range,
np.median(particle_filter_output.compartments[:, :, 0, 3], axis=0).squeeze(),
color="black",
label="median of PF",
)
axes[2, 0].legend()
axes[2, 1].set_xlabel("Days")
axes[2, 1].set_title("Fit to the $\\beta$")
axes[2, 1].fill_between(
sim_date_range,
np.exp(
np.percentile(
particle_filter_output.estimated_params[NamePattern.of("log_beta")],
2.5,
axis=0,
)
).squeeze(),
np.exp(
np.percentile(
particle_filter_output.estimated_params[NamePattern.of("log_beta")],
97.5,
axis=0,
)
).squeeze(),
label="95% CI",
alpha=0.3,
)
axes[2, 1].plot(sim_date_range, true_beta, color="red", label="Data points")
axes[2, 1].plot(
sim_date_range,
np.exp(
np.median(
particle_filter_output.estimated_params[NamePattern.of("log_beta")], axis=0
)
).squeeze(),
color="black",
label="median of PF",
)
axes[2, 1].legend()
fig.tight_layout()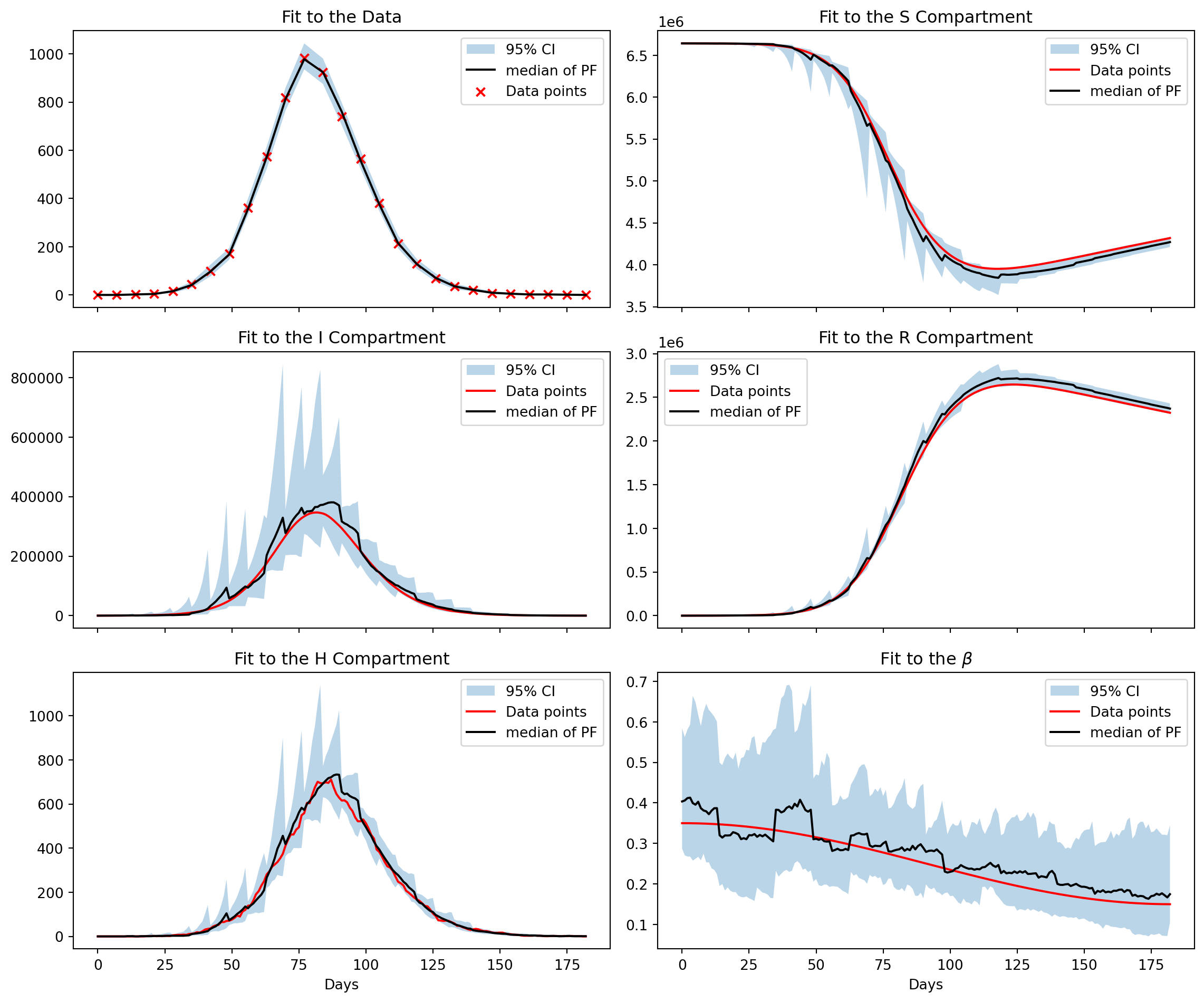
Notes
We see a strong fit to the data and that the algorithm has captured the downwards trajectory of \(\beta\). While the median of the state variables hews closely to their true values we also observe a sawtooth effect in the 95% credible intervals. This is due to the sparsity of the data, weekly conditioning leads to sharp adjustments to the posterior distribution.