epymorph: Spatially-explicit modeling of infectious disease as a Python library

The epymorph package is the product of the EpiMoRPH (Epidemiological Modeling Resources for Public Health) project, and aims to provide a simplified framework for completing the full lifecycle of a spatial modeling experiment. epymorph streamlines methods for building, simulating, and fitting, metapopulation models of infectious pathogens.
With its focus on automation, simple python syntax, and conceptual simplicity, the epymorph package is easily accessible to beginning modelers, while also providing sophisticated features for rapid design and execution of complex modeling experiments by highly experienced modelers. Specific aims include dramatic streamlining of model building speed, increased model transparency, automated fitting of models to observed data, and easy transportability of models across temporal and geographic scenarios. The epymorph conceptual model, in partcular, is designed to serve as a common basis for describing, sharing, and disseminating spatially explicit metapopulation models within the modeling community.
In the remainder of this page, we briefly introduce the epymorph conceptual model, and then highlight some of the powerful modeling capabilities it enables in our package.
GPM: a uniform framework for metapopulation modeling
The EpiMorph project — and the resulting epymorph package presented here — is fundamentally motivated by a desire to dramatically streamline the construction, fitting and evaluation of spatially explicit metapopulation models; and the ability to share, analyze and test evolving metapopulation modeling approaches within the modeling community. See the About page for more information about the motivations behind EpiMoRPH. In a nutshell, the fundamental obstacle to rapid construction, sharing, and analysis of generality is that, currently, metapopulation models are implemented as custom, hand-built, statistic programs; implemented in a variety of languages and individually architected based on the unique preferences of individual modelers.
Epymorph introduces a simple conceptual framework called Geography-Populations-Movement, or GPM, as a shared, uniform basis for all metapopulation models. The GPM concept essentially declares that all spatially-explicit metapopulation models can be expressed in terms of three distinct, interacting sub-models or components:
An Intra-Population Model, or IPM. This sub-model describes the pathogen dynamics within population nodes (e.g. states, cities, census blocks, etc.) within the model. An IPM is just a standard compartment model, with the usual rate equations, initial conditions, and parameters. Parameters may be static, time-dependent, or time- and node-dependent; parameters and initial conditions may be explictly specified values/arrays, or could refer abstractly to values (e.g. populations, distances, temperatures, etc.) from the geographic location the model is running in.
A Movement Model, or MM. This sub-model specifically describes the movement pattern of individuals between population nodes in the spatially-explicit model. The MM essentially gives a statistical description of the likelihood of movement between each population node for individuals within compartments within those nodes. The MM can be a constant value, time-dependent, or time/node-dependent; it can be provided by the user or drawn from large databases (e.g. U.S. Census, LODES8, etc.).
A Geographic model, or GEO. In many cases, a modeler’s theory of how a pathogen spreads, either within a population node or through movement between nodes (IPM or MM), depends on or refers to the geography that the paired IPM+MM is running in, i.e., where and when the model is run. The GEO can be seen as the enclosing context of the modeling experiment, providing all modeling data specified as relevant in the IPM or MM, e.g., population node sizes, distances between population nodes, travel patterns of individuals, temperature, humidity — whatever “local values” the modeler has brought into play in the design of the IPM or MM. The GEO also include a timeframe for the modeling experiment, allowing epymorph to automatically fetch many of the required data from relevant public databases for that time period.
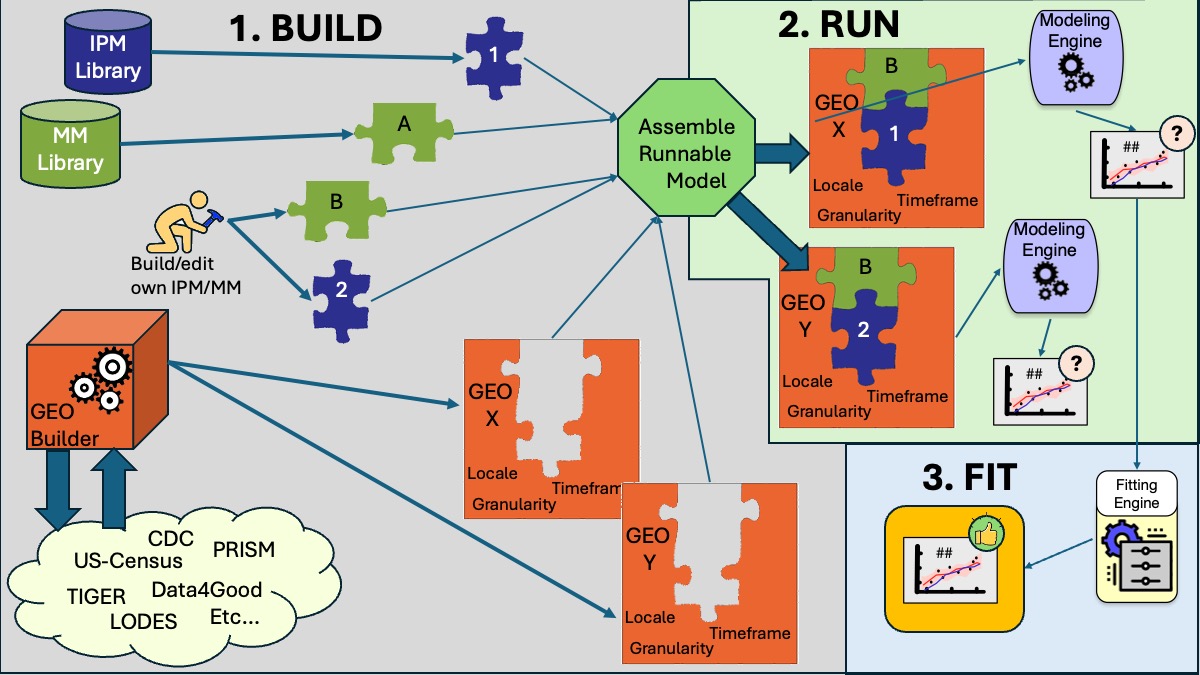
As suggested by Figure 1, the power of the GPM concept lies in the clear, modular separation of infection, movement, and geo-temporal specification of any disease model. With a common framework and component interfaces, a modeler can freely “snap together” any desired IPM with a any desired MM, and then run those within any selected GEO and timeframe. Moreover, modelers can easily share, compare, and evaluate IPMs and MMs within the modeling community; a simple uniform syntax for IPMs, MMs, and GEOs make modeling paramters and assumptions accessible and clear to all. Finally, a growing library of standard or common IPMs and MMs can develop, allowing modelers (including beginners or learners) to build and explore relatively sophisticated disease models just by snapping together pre-built IPMs and MMs, then running the model in any desired timeframe. Finally, GPM-based modeling experiments can be easily exported, saved, and shared with others, providing a powerful, clear, common basis for critical scientific discussion about modeling approaches.
Building on this quick intro to modeling using the EpiMoRPH GPM approach, let’s take a quick look at some core features of the epymorph package!
Epymorph core Features: Build and explore spatial models fast!
Leveraging the GPM concept, epymorph is able to standardize and largely automate significant parts of the workflow for creating, simulating, vizualizing, and fitting spatial epidemiological models. Rather than coding all infection, movement and local geographic aspects of a model by hand and from scratch every time, modelers can re-use or modify standard IPM or MM components, inventing and exploring only those aspects of a model they are focused on. Although modelers can still easily build IPMs and MMs from scratch, a more common pattern in epymorph is to start with a “similar” IPM or MM developed by epymorph team members or other modelers, then modify or extend that element in some way to explore a new hypothesis of disease dynamics. Such modified or novel IPM and MM elements can (optionally) be shared with the community, greatly accelerating model exploration and refinement for a given pathogen.
The epymorph package is packed with powerful features to support and streamline this modular, snap-together approach to metapopulation modeling enabled by our GPM concept. Let’s briefly look at just a few:
- Pre-built catalogs of mix-and-match IPM and MM components The standardized, modular design of epymorph means that our pre-built library of IPMs and MMs are cross-compatible; users can quickly select, modify, and click together different movement patterns (MMs) with different compartmental models (IPMs) to suit their specific interests or to test certain hypotheses (e.g., how different movement styles affect local and regional patterns of disease). Once a promising IPM+MM combination is discovered, it can easily be situated, run, and evaluated in arbitrary geographic locations or timeframes — all within hours rather than after days or weeks of effort.
Within this workflow, locating and extracting the actual data needed to drive a modeling experiment for a partiular timeframe or geographic location is often the most arduous and time-consuming aspect of the process. Let’s look next at how epymorph streamlines this!
A powerful GEO subsystem to automate access to modeling data, for any location, timeframe, or granularity A powerful GEO subsystem not only allows modelers to easily specify the geographic scope, granularity, and targeted timeframe for a modeling experiment, but also provides a growing set of powerful data access and retrieveal tools, called Abstract Data Retrieval and Integration Objects (ADRIOS), that are able to automatically extract and retrieve data needed by the model from publicly accessible databases. Each ADRIO is essentially “an expert” on its public data source; modelers have no need to learn the complex data models and APIs of those data sources, they simply reference the desired data in their model specification and epymorph takes care of the rest.
Data currently made accessible via epymorph ADRIOS at state, county and (where available) census block group scales include local popuplations and demographic characteristics (distribution based on age, economic factors, etc.), commuter data, temperature, precipitation, relative humidity and more.Built-in time-series and geo-spatial plotting functions Another advantage of the standardization and modularity built into epymorph is that simulation outputs can all be captured in a standard, uniform output object. This allows epymorph to provide a powerful, built-in data analysis and visualization module that allows users to very quickly examine and visualize modeling output. This allows inexperienced modelers to easily visualize outputs without first building significant expertise with complex general purpose graphing and visualization tools. Expert modelers benefit as well, able to take a quick look at key model outputs during their interactive model development and refinement process — and then download detailed final model output in standard formats for further customized graphing and visualization. Easy built-in tools include a wide variety of selectable time-series line plots (e.g., infections over time per model node), and configurable static maps (e.g., choropleth maps plotting summary statistics per GEO node).
Features for Advanced Modelers
The features outlined above are specifically designed to make the process of building, exploring, and evaluating meta-population models more easily accessible and understandable for students and other non-experts in modeling. At the same time, the modularity and automation provided by epymorph greatly streamlines expert model development as well. Here are just a few of the advanced features epymorph provides for modeling experts:
Built-in parameter estimation with particle filtering, supported by real surveillance data sets Advanced users with experience in model-fitting can leverage our built-in model fitting engine to fit epymorph models to real or simulated surveillance data. This major advance allows modelers to quickly test the quality of hindcasts produced from different IPMs and MMs fit to real data sets, enabling advanced hypothesis testing and the design of forecasting pipelines. The epymorph model-fitting engine provides a growing number of flexible and powerful particle filtering algorithms for both time-varying and static parameter estimation, while our GEO subsystem can automatically extract and retrieve real surveillance data (e.g., hospitalization or mortality counts) for influenza, SARS-CoV-2, and RSV from public CDC databases. Users may also provide their own private surveillance data sets (e.g., simulated or unreleased) for use by our fitting engine. Parameters can be estimated per node in the GEO Scope, using built-in localization techniques, which provides a high-quality understanding of spatial heterogeneities in transmission and disease processes.
Rapidly create and customize advanced intrapopulation models As noted, epymorph provides a library of relatively simple, standard compartment models out of the box, including models that capture hospitalization, deaths, vaccinations and other common compartments. Using our growing set of sample “vignettes”, user documentation, and epymorph API documentation, advanced users can quickly learn to “program in epymorph”, i.e., to modify or extend existing IPMs to their purpose, or create completely novel IPMs from scratch — all in just a page or two of easily readable epymorph code, using our straightforward and familiar syntax. Users can also use our multi-strata modeling functions to easily create sophisticated composite models with sub-strata based on age-classes, risk-classes, or host species (e.g., vector-borne or zoonotic diseases). A separate IPM can be selected (or modified/built) and attached to each stratum, then combined into a composite model by linking influences or compartments between strata. Similarly, a separate movement model (MM, see next bullet) can be selected (or modified/built) and connected to each stratum, e.g., to model different movement dynamics per age-class, or different movement dynamics per species in a multi-species model.
Flexibly define complex, multi-faceted, customized movement models Movement of individuals within the model geography is critical for modeling the dynamics of disease spread, capturing how infected individuals or vector hosts move the population between nodes (cities, study sectors, subpopulatiions, etc.). Just as with IPMs, epymorph users combine and modify our pre-built movement models, or they can create their own movement models to embody novel theories of host movement, or that leverage actual real-time datasets of movement data in public datasets (see GEO discussion above), or directly available to the user. Our carefully developed MM specification syntax allows movement models to capture an enormous variety of potential movement patterns including:
- Migration, commuter movements, and custom-triggered movement.
- Overlapping movement dynamics, e.g., weekday versus weekend movement, or commuter movement that happens multiple times per day.
- Seasonal migrations, or movements on particular holidays.
- Movement drawn from public databases (e.g. airline flight data).
- Movement described by custom user-defined functions that create custom movement matrices, for instance through customized gravity kernel functions that leverage data from each node in the GEO Scope.
The epymorph movement engine effectively “compiles” all of movement dynamics specified by the user, combining all the effects into a coherent, combined movement matrix used to drive inter-node movements for each node and compartment for each model time step.
- Specifying custom geographies Epymorph is natively designed to operate on the geographic delineations defined by the U.S. Census (state, county, census block group, etc.); most of the ADRIOs in our GEO subsystem extract data from public databases, since these databases typically are based around these standard delineations. Advanced users can, however, define their own model nodes based on their specific needs. For example, some state health departments report infection or hospitalization data by zip codes, which are not congruent with US Census geographic delineations; similarly, study areas for animal disease data collection may not fit nicely to Census delineations. In such cases, epymorph provides tools to allow advanced users to specify custom geographic scopes (node spaces) — yet still fetch other data from standard public data sources, using custom functions to adapt that data to their unique geographic scope. Alternatively, users can use epymorph to import their own datasets from CSV files; epymorph will help to ensure the imported data is formatted and shaped correctly to match the selected modeling granularity, geography, and node structure. The plug-n-play modularity of the GPM concept underlying epymorph means that IPMs and MMs simply don’t care what geographies they operate within; once a coherent node space and modeling timeframe has been constructed, any IPM or MM can be run in that GEO scope.